Results reported in the paper KAnalyze: A Fast Versatile Pipelined K-mer Toolkit in Bioinformatics about DSK don’t seem to fit the reality.
Taking the same datasets, we run DSK and Kanalyze on our machines.
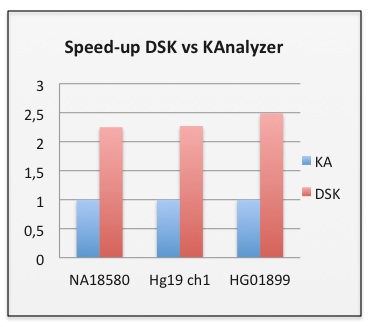
The following table summarizes our experiments.
| KAnalyze | DSK | ||
| NA18580 | 1.5 M reads – 453 Mbp | 201 s | 89 s |
| Hg19 ch1 | 243 Mbp | 173 s | 76 s |
| HG01899 | 988 M reads – 72 Gbp | 11h 1 m 57s | 4h 28 m 23 s |
times are ‘real’ time reported by ‘time’ command
The test machine is an intel E5-2670 @ 2.5 GHz with 64GB ram, running fedora 20. The disk used is a 2TB HDD ( seagate barracuda ST2000DM001)
Both softwares were run with 2 threads, 2 GB max of ram (default parameters for kanalyze), and k=31.
Disk usage
For the HG01899 dataset, kanalyze used 352 GB of disk space to store its temporary files, while DSK required only a maximum of 75 GB of temp space.
In the KAnalyze paper, authors claim DSK kmers count did not agree with other software results. While in fact, DSK and Kanalyze outputs exactly the same results when running kanalyze with the -rcanonical option (equivalent of -C option in jellyfish, and default in dsk).
Parse/postprocess
In their KAnalzye paper, authors add to the dsk run time a ‘parse’ time to convert the dsk result file (stored in binary) to a human-readable ASCII format. While dsk 1.5280 indeed included a python script to perform such conversion, an ascii result file is only needed for manual inspection or debugging. Real world usage of DSK inside an assembler or an aligner will work directly on the binary results file, which is much more compact and more efficient to read. Therefore a ‘parse’ conversion time has no reason to be included in total runtime.
Furthermore, since October 2013, DSK releases include an efficient parser program written in C++.
Software Version used
KAnalyze 0.9.3.
DSK included in the GATB library 1.0.0 . (no major algorithms changes or optimizations have been done since dsk 1.5280 version)


